Associate Professor in Biology
Education
Ph.D., Massachusetts Institute of Technology
B.A., Amherst College
About
I teach Biochemistry, Genetics, Seminars in Biology and Biochemistry, and two research-based courses: Research Experience in Biology and Experimental Biochemistry. I am also the co-coordinator for the Biochemistry program, along with Professor Hilary Gaudet in the Chemistry Department. I am especially interested in helping students acquire the skills to become confident scientists, from designing and executing research projects to learning how to read, interpret, and contribute to the scientific literature. Beyond the lab bench, I am committed to developing effective strategies to combat racism in biology and pre-health education.
Courses taught
BIO112 – Cells and Genes
BIO211 – Genetics
BIO200 – Research in Regenerative Biology
BIO305 – Biochemistry
BIO401 – Biology Senior Seminar
BIO405 – Biochemistry Senior Seminar
FSEM101 – Cancer: Exploring the Enemy Within
Research Interests
- Growth regulation in vertebrates
- Regenerative biology
- Anti-racism in science education
Publications
Peer-reviewed scientific publications
Daane, J. M., Blum, N., Lanni, J., Boldt, H., Iovine, M. K., Higdon, C. W., Johnson, S. L., Lovejoy, N. R., & Harris, M. P. (2021). Modulation of bioelectric cues in the evolution of flying fishes. Current Biology : CB, 31(22), 5052–5061.e8. https://doi.org/10.1016/j.cub.2021.08.054
Harris, M.P., Daane, J.M. and Lanni, J. (2021). Through veiled mirrors: Fish fins giving insight into size regulation. WIRES Developmental Biology . https://doi.org/10.1002/wdev.381
Lanni, J.S., Peal, D., Ekstrom, L., Chen, H.*, Stanclift, C.*, Bowen, M., Kahle, K., Harris, M. (2019) Integrated K+ channel and K+Cl- cotransporter functions are required for the coordination of size and proportion during development. * denotes undergraduate author. Developmental Biology 456(2), 164-178. https://doi.org/10.1016/j.ydbio.2019.08.016
Daane, J., Lanni, J., Rothenberg, I., Seebohm, G., Higdon, C., Johnson, S., & Harris, M. (2018). Bioelectric-calcineurin signaling module regulates allometric growth and size of the zebrafish fin. Scientific Reports 8, Article number:10391 https://doi.org/10.1038/s41598-018-28450-6
Opinion and commentary
Lanni, J. (2022, Nov. 9). Bridging the gap: A biologist considers the legacies of slavery. Legacies of American Slavery. https://legaciesofslavery.net/2022/11/09/bridging-the-gap-a-biologist-considers-the-legacies-of-slavery/
Lanni, J. (2021). Why I teach my students about scientific failure. Science, 374(6575), 1642.
https://www.science.org/content/article/why-i-teach-my-students-about-scientific-failure
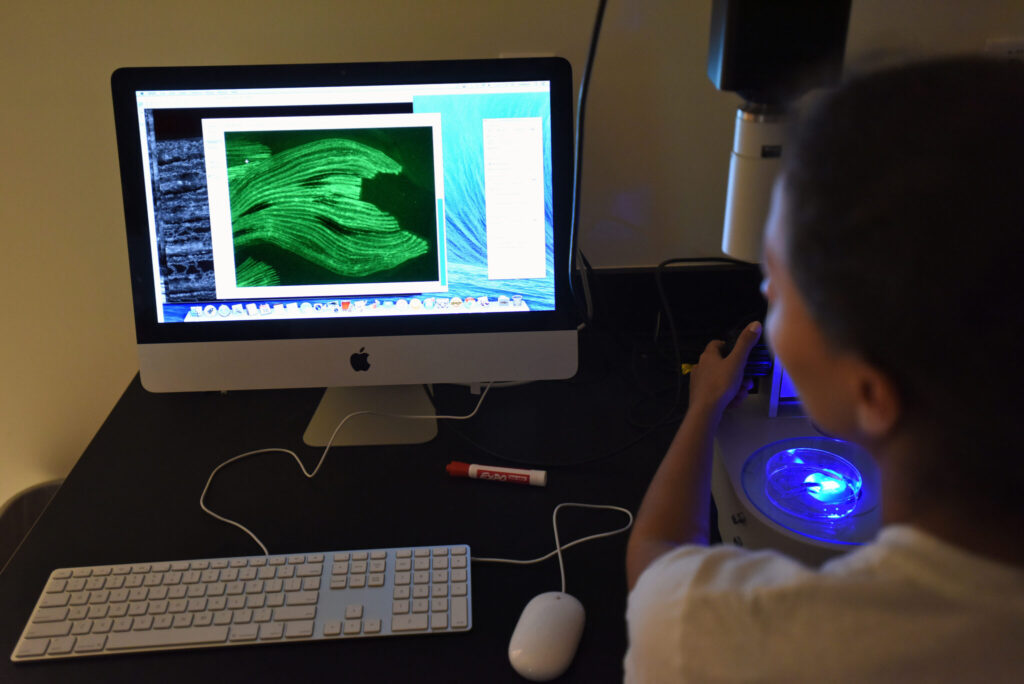

Student Projects
William Gan and Jeanne Specht, 2021: Quantitative analysis of sprouting angiogenesis in the zebrafish fin.
Haining Chen, 2019: Genotypic and phenotypic analysis of unique Crispr-Cas induced mutations in the KCC4a gene.
Michelle Laverriere, 2018: Quantitative measurement of gene expression in wild-type, mutant, and regenerating zebrafish fins using qPCR.
Liam McCafferty, 2017: RNAseq analysis of gene expression in wild-type and mutant zebrafish fin tissue.
Caroline Stanclift, 2017: Immunohistochemical detection of Kcc4 protein in zebrafish tissues.
Katie Henrikson, 2016 and Kevin (Ao) Shi, 2017: Phenotypic analysis of the vasculature and pigment during fin regeneration in zebrafish strains with activated potassium channels.
Ethan Fitzgerald, 2016: Determining the pattern and timing of KCC4a expression in wild-type and mutant zebrafish strains. (Also worked on phenotypic analysis)
Julia Jennings, 2015: Assaying the lateral line phenotype in zebrafish strains with potassium channel mutations. (Also worked on phenotypic analysis)


Research Interests
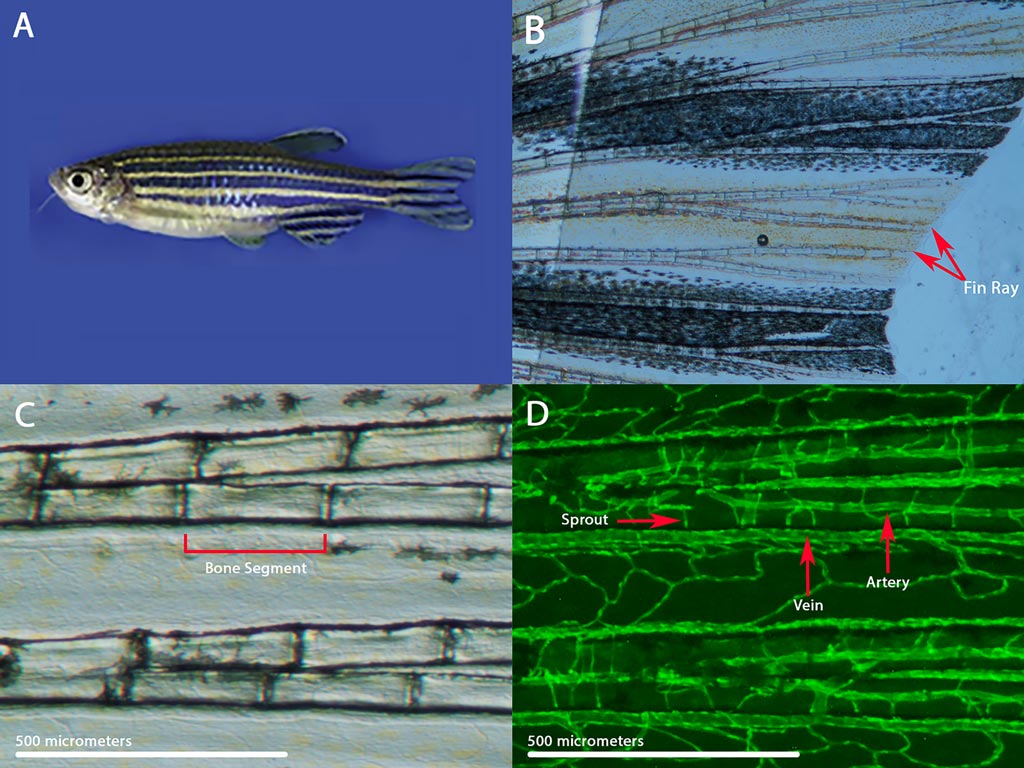
My laboratory research utilizes the zebrafish model system to explore the regulation of proportional growth in vertebrates. During normal development, growth is integrated such that relative sizes among structures and tissues are specified and maintained. My laboratory is studying zebrafish mutant strains with fins that grow to over twice the normal length. These long-finned zebrafish are notable in that they maintain patterned overgrowth, distinct from the kind of aberrant proliferation seen in cancer and overgrowth disorders.
Interestingly, a mutation in any one of several different genes that encode potassium transport proteins can cause the long-finned phenotype. This finding suggests a functional connection between potassium ion homeostasis and vertebrate growth regulation. Our current work is focused on elucidating the changes in gene and protein expression that differentiate long-finned fish from wild-type fish, and then using genetic and biochemical tools to determine which of these changes are causally linked to fin overgrowth.
Zebrafish also possess the remarkable ability to fully regenerate their fins within two weeks of amputation. Thus, understanding the growth pathways that are activated in our long-finned fish may lend insight into the molecular underpinnings of tissue regeneration. Also, because zebrafish share many of their genes with humans, our work may identify a conserved role for potassium ion homeostasis in vertebrate growth regulation.
More recently, I have started to pursue research in STEM education. My particular focus is developing effective strategies to combat racism in biology and pre-health education. My research is on how to effectively integrate discussions of race into our core Biology and Nursing courses, in order to address racialized misconceptions in anatomy, physiology and genetics.
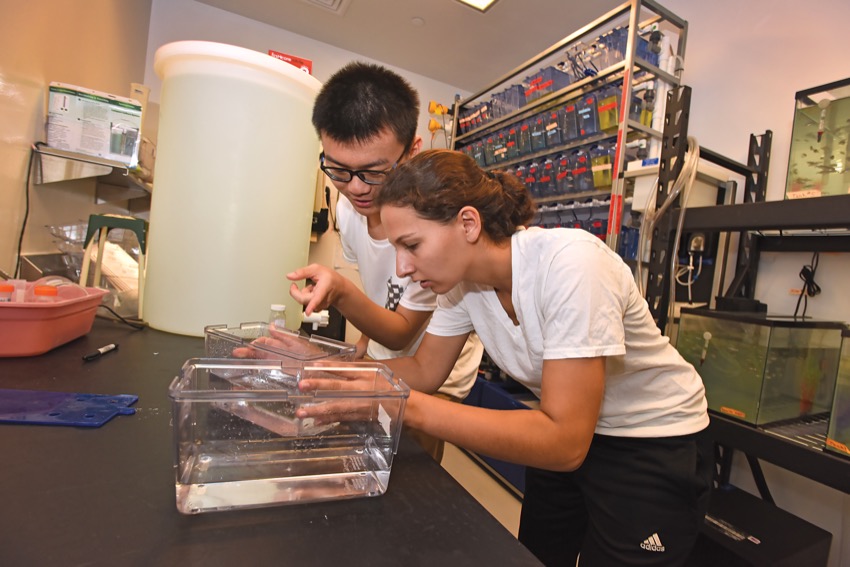

Ao “Kevin” Shi ’17 and Kathryn Henrikson ’16 in the fish room
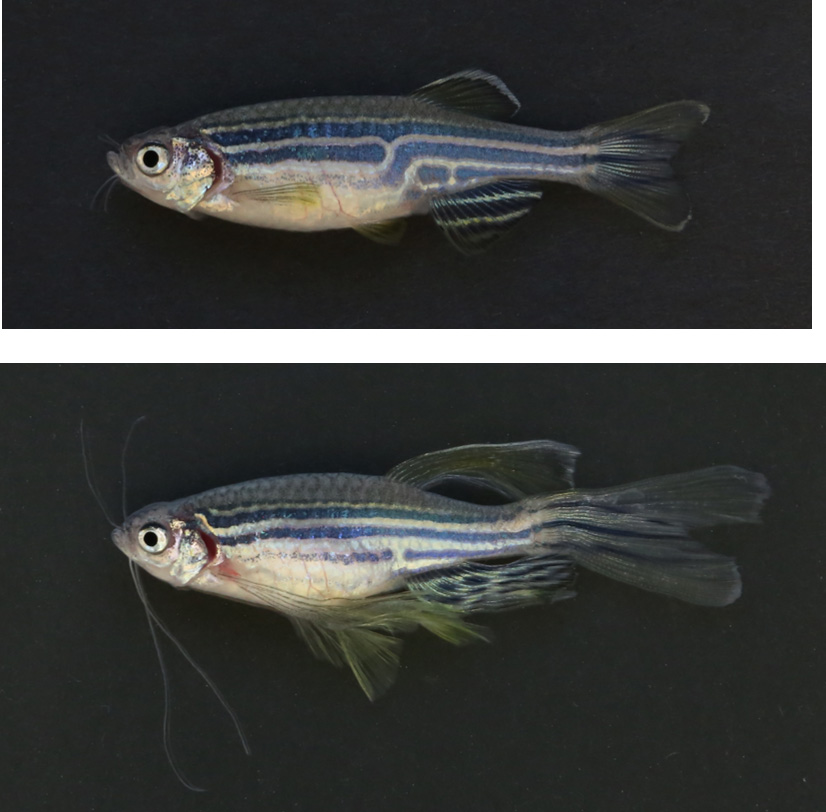

Wildtype and longfinned zebrafish (Danio rerio)
